
Université Paris-Saclay
Biography
My research aims to address fundamental issues in self-organizing biological matter at the microscopic scale using experimental and theoretical concepts derived from soft condensed matter physics. In particular, I focus on the high-resolution structure of ordered and weakly disordered systems, as well as on the nonequilibrium dynamics of assembly and self-organization phenomena. I exploit state-of-the-art techniques including cryotransmission electron microscopy and time-resolved radiation scattering with large-scale facilies, and develop models that account for the observed complex behaviors.
Interests
- Biological self-assembly and self-organization
- Viruses
- DNA condensates
- Small-angle scattering
- Microfluidics
Education
- Accreditation to supervise research (2013), Université Paris-Sud
- PhD in physics (2002), Université Aix-Marseille I
- MSc. in physics (1998), Université Paris-Sud
- MEng. in electrical engineering (1998), Ecole Supérieure d’Electricité (Supélec)
Statistical data analysis
Master 2 Systèmes Biologiques et Concepts Physiques
Doctoral school EDPIF Physique en Ile-de-France
Five or ten 3-hours sessions each split into half lecture and half practicals in Python.
Research in physics requires the acquisition, analysis and interpretation of experimental measurements in order to verify an hypothesis and/or devise a theoretical model. The great variability inherent to many physical systems makes it essential to take into account the uncertainties and to use powerful statistical tools available today via Python. Each session consists of a first theoretical part that presents the fundamental concepts of statistical data analysis, illustrated in the second part of the course by practical work in Python through concrete situations encountered in physics and biophysics. The main topics covered include a reminder of probabilities and distributions, statistical tests (chi2, Student, p-value), handling experimental uncertainties (confidence intervals), linear and non-linear adjustments (least squares, covariance matrix, correlations), and some advanced notions (bayesian inference, statistical learning).
Small-angle X-ray scattering and colloidal systems
Ecole Supérieure de Physique et de Chimie Industrielle de la Ville de Paris (ESPCI)
Two sessions of 2 hours each.
This course is an introduction to small-angle X-ray scattering applied to dilute and crystallized colloidal systems. It gives the fundamentals of X-ray scattering by one electron, then the interferences obtained in the presence of two electrons and the generalization with a continuous electron density. The main results for a solution of spherical particles are deduced analytically. Actual scattering data of colloidal suspensions are analyzed thus providing the methodology for characterizing nanoparticles and liquid crystals at the microscopic scale.
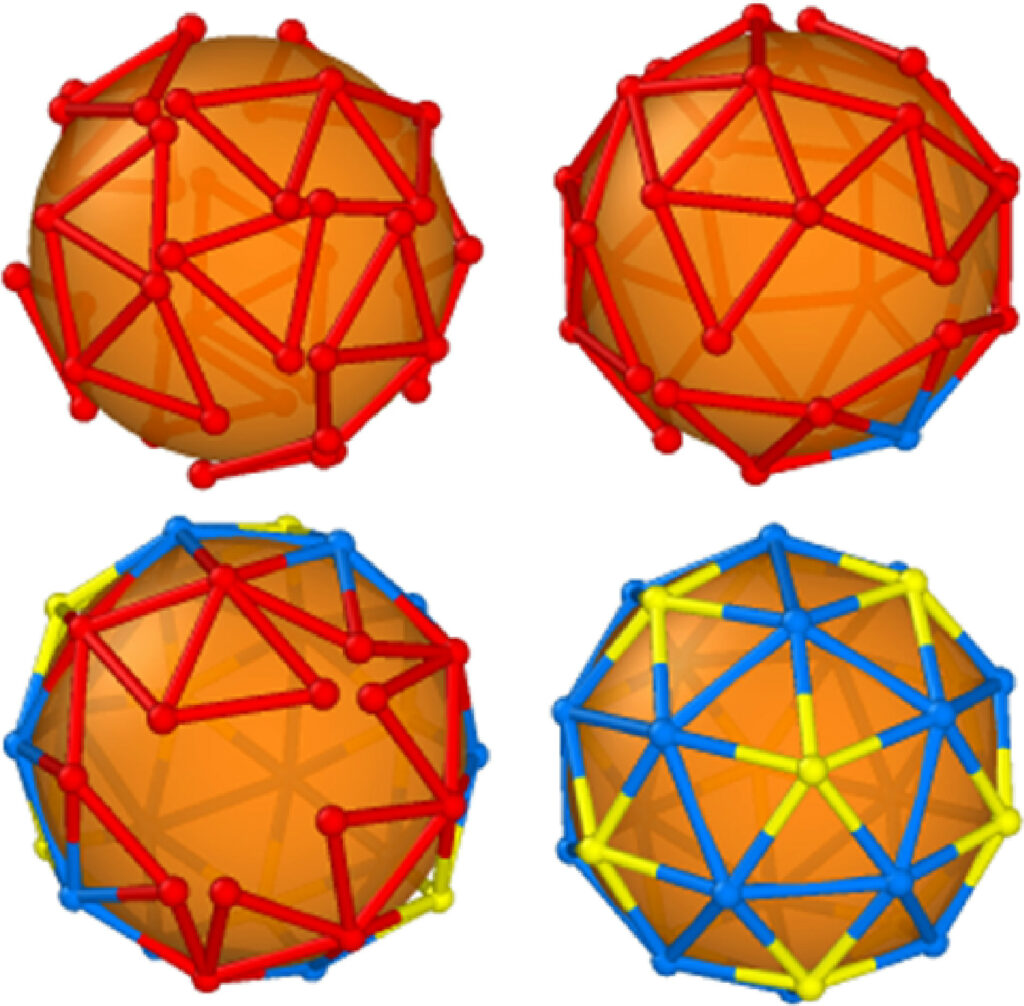
Virus self-assembly in complex environments
Viruses are amazing biological agents in which hundreds of molecular blocks integrate with atomic precision into the final structure. Their regularity is all the more remarkable because for many viruses it occurs spontaneously, in an efficient self-assembly process, whether in the host cell or in a test tube. Many viruses are assembled in a crowded intracellular environment, with a low error rate, despite numerous nonspecific interactions with cellular components. Our objective is to decipher the self-assembly pathways of icosahedral viruses in complex, cell-like environments, by setting up a cross-disciplinary strategy that combines experimental physics and quantitative biology with an advanced soft-matter theoretical framework.
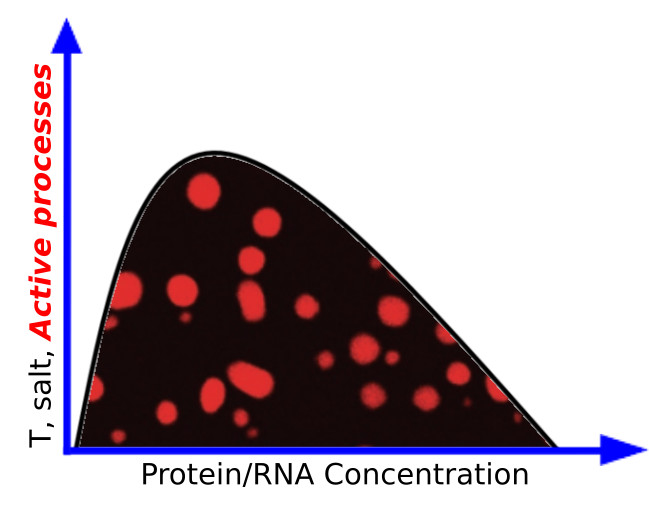
Out-of-equilibrium cellular droplets
Living cells are organized into distinct subcompartments to facilitate spatiotemporal regulation of biological reactions. While this organization is traditionally thought to rely on membrane-delimited organelles, such as the nucleus and mitochondria, it also involves membrane-less assemblies of proteins and nucleic acids that are kept together only by weak, dynamical interactions between their components. In the last years groundbreaking studies suggested that these cellular assemblies have liquid-like physical properties and are formed via liquid-liquid phase separation (LLPS) processes. We plan to rely on state-of-the-art techniques in soft matter physics and on an advanced molecular modeling framework for unveiling general features of cellular LLPS in equilibrium and nonequilibrium conditions
and linking them back to their molecular origin.
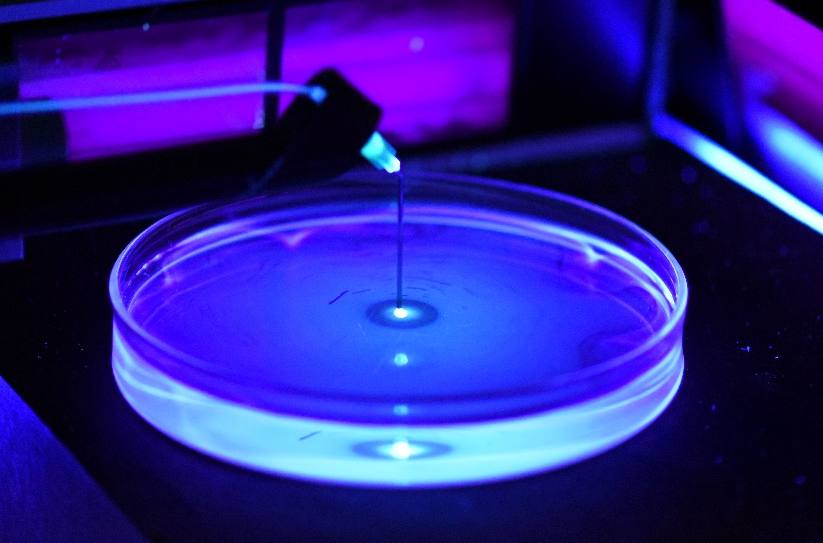
Marangoni flows
The Marangoni effect is a spectacular manifestation of the interfacial forces driven by surfactants at the air-water interface. It stems from the gradients of surface tension due to the inhomogeneity of the surfactant density across the interface, which subsequently induce the
motion of water over a thin layer. This project aims at elucidating the flows generated in the bulk by visualizing them with a fluorescent probe.
Contact
- guillaume.tresset@universite-paris-saclay.fr
- +33 (0) 1 69 15 53 60
- Laboratoire de Physique des Solides
Université Paris-Saclay
rue Nicolas Appert
Building 510
91405 Orsay Cedex
France
Laboratoire de Physique des Solides, Université Paris-Saclay, CNRS (Orsay, France)
Jan 2021 – present: Deputy director
Oct 2019 – present: CNRS research director
Jul 2019 – Dec 2020: Head of the “Self-Assembled Biological Objects” team
Oct 2008 – Oct 2019: CNRS research associate
Institute of Bioengineering and Nanotechnology, A*STAR (Republic of Singapore)
Jan 2005 – Sep 2008: Research scientist
Laboratory for Integrated Micro Mechatronic Systems, The University of Tokyo (Japan)
Nov 2002 – Dec 2004: Postdoctoral fellow
Département de Recherches pour la Fusion Contrôlée, CEA (Saint-Paul-lèz-Durance, France)
Oct 1999 – Sep 2002: Doctoral candidate