Liquid crystalline nucleosomes in vitro
We design in vitro model systems of condensed chromatin, explore their physico-chemical properties and establish phase diagrams.
Nucleosome core particles are formed by 145–147 bp DNA wrapped into 1.65 turn of a left-handed superhelix around a histone octamer (two copies of H2A, H2B, H3 and H4). We explore their self assembly, varying different parameters : (i) the osmotic pressure, (ii) monovalent salt concentration, (iii) the presence of di or multivalent cations. Under physico-chemical conditions that reproduce those found in the living cell, nucleosomes form multiple amorphous, liquid crystalline and crystalline phases that can be related to the poly-ampholyte and geometric characteristics of the particle. All phases also present complex mesoscopic chiral organisations with multiple levels of helicity.

Phase diagram of nucleosome core particles in the presence of monovalent counterions.
Nucleosome conformational variability in vitro and in situ
Using cryo electron microscopy of vitreous sections we analyse the conformation of native nucleosomes in vitro (purified particles solubilised at physiologically relevant concentrations) and in situ within interphase nuclei. We demonstrate a salt-dependent transition, with a high salt compact conformation resembling the canonical nucleosome, and an open low salt one, closer to nuclear nucleosomes.

Local organisation of the eukaryotic chromosome: the CRYOCHROM project
Eukaryotic chromosomes are complex polymorphic and dynamic objects. Centimeters to meters of DNA condense into a micron-scale nucleus though multiple levels of structural organisation, while interacting with a multitude of factors driving functional compartmentalization. Their basic beads-on-string nucleosome structure plays a key role in major cellular processes from regulation of gene expression and epigenetics to DNA replication and repair, cell differentiation and aging.
Using cryo electron microscopy and tomography of vitreous sections,we unveil the conformation and local order of nucleosomes and chromatin in the nuclear context, in particular in the interphase nucleus. Following an integrated structural approach we aim to relate chromatin structure and functional compartmentalization and benchmark chromatin in active and repressed states.
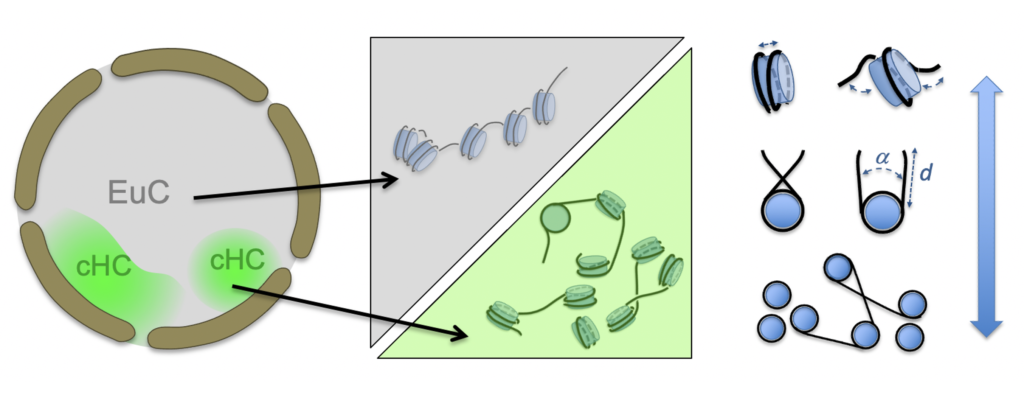
CRYOCHROM is a multidisciplinary collaborative project funded by the ANR, in partnership with M. Eltsov (IGBMC, Illkirch), S. Jonic (IMPMC, Sorbonne Université) and J.-M. Victor (LPTMC, Sorbonne Université).
Contact
Amélie Leforestier: amelie.leforestier@universite-paris-saclay.fr
Related publications
- Harastani, M., Eltsov, M., Leforestier, A., & Jonic, S. (2022) TomoFlow: Analysis of continuous conformational variability of macromolecules in cryogenic subtomograms based on 3D dense optical flow. Journal of molecular biology, 434(2), 167381. doi.org/10.1016/j.jmb.2021.167381
- Harastani, M., Eltsov, M., Leforestier, A., & Jonic, S. (2021). HEMNMA-3D: Cryo Electron Tomography Method Based on Normal Mode Analysis to Study Continuous Conformational Variability of Macromolecular Complexes. Frontiers in molecular biosciences, 8. doi: 10.3389/fmolb.2021.663121
- Eltsov, M., Grewe, D., Lemercier, N., Frangakis, A., Livolant, F., & Leforestier, A. (2018). Nucleosome conformational variability in solution and in interphase nuclei evidenced by cryo-electron microscopy of vitreous sections. Nucleic acids research, 46(17), 9189-9200. doi.org/10.1093/nar/gky670
- Leforestier, A., Bertin, A., Dubochet, J., Richter, K., Blanc, N. S., & Livolant, F. (2008). Expression of chirality in columnar hexagonal phases or DNA and nucleosomes. Comptes Rendus Chimie, 11(3), 229-244. doi.org/10.1016/j.crci.2007.09.008
- Livolant, F., Mangenot, S., Leforestier, A., Bertin, A., Frutos, M. D., Raspaud, E., & Durand, D. (2006). Are liquid crystalline properties of nucleosomes involved in chromosome structure and dynamics?. Philosophical Transactions of the Royal Society A: Mathematical, Physical and Engineering Sciences, 364(1847), 2615-2633.
- Bertin, A., Leforestier, A., Durand, D., & Livolant, F. (2004). Role of histone tails in the conformation and interactions of nucleosome core particles. Biochemistry, 43(16), 4773-4780.
- Mangenot, S., Leforestier, A., Durand, D., & Livolant, F. (2003). Phase diagram of nucleosome core particles. Journal of molecular biology, 333(5), 907-916. doi.org/10.1016/j.jmb.2003.09.015
- Mangenot, S., Leforestier, A., Durand, D., & Livolant, F. (2003). X-ray diffraction characterization of the dense phases formed by nucleosome core particles. Biophysical journal, 84(4), 2570-2584.
- Leforestier, A., Dubochet, J., & Livolant, F. (2001). Bilayers of nucleosome core particles. Biophysical journal, 81(4), 2414-2421. doi.org/10.1016/S0006-3495(01)75888-2
- De Frutos, M., Raspaud, E., Leforestier, A., & Livolant, F. (2001). Aggregation of nucleosomes by divalent cations. Biophysical journal, 81(2), 1127-1132.
- Livolant, F., & Leforestier, A. (2000). Chiral discotic columnar germs of nucleosome core particles. Biophysical journal, 78(5), 2716-2729. doi.org/10.1016/S0006-3495(00)76816-0
- Raspaud, E., Chaperon, I., Leforestier, A., & Livolant, F. (1999). Spermine-induced aggregation of DNA, nucleosome, and chromatin. Biophysical journal, 77(3), 1547-1555.
- Leforestier, A., Fudaley, S., & Livolant, F. (1999). Spermidine-induced aggregation of nucleosome core particles: evidence for multiple liquid crystalline phases. Journal of molecular biology, 290(2), 481-494.
- Leforestier, A., & Livolant, F. (1997). Liquid crystalline ordering of nucleosome core particles under macromolecular crowding conditions: evidence for a discotic columnar hexagonal phase. Biophysical journal, 73(4), 1771-1776.